QCElemental API
qcelemental Package
Functions
|
Returns True if two integers, strings, booleans, or integer arrays are element-wise equal. |
|
Recursively compares nested structures such as dictionaries and lists. |
|
Returns True if two floats or float arrays are element-wise equal within a tolerance. |
Classes
|
Error called for problems with syntax input file. |
|
Covalent radii sets. |
|
Error when dataset incomplete and otherwise valid query can't be fulfilled. |
Facilitates the storage of quantum chemical results by labeling them with basic metadata. |
|
|
Error called when a molparse.from_string contains unparsable lines. |
|
Error when element or nuclide can't be identified. |
|
CODATA physical constants set from NIST. |
|
Error called for problems with syntax input file. |
|
Van der Waals radii sets. |
Variables
CODATA physical constants set from NIST. |
|
Covalent radii sets. |
|
Nuclear and mass data about chemical elements from NIST. |
|
Van der Waals radii sets. |
Class Inheritance Diagram

qcelemental.molparse Package
Functions
|
Take (nat, ?) array-like arrays and return with atoms arranged by (nfr, ?) frag_pattern. |
|
Compose a Molecule dict from unvalidated arrays and variables, returning dict. |
|
Compose a Molecule dict from unvalidated arrays and variables in multiple domains. |
|
Construct molecule dictionary representation from non-Psi4 schema. |
|
Construct a molecule dictionary from any recognized string format. |
|
Separate molecule nucleus string into fields. |
|
Forms consistent set of nucleus descriptors from all information from arguments, supplemented by the periodic table. |
|
Translate molparse internal Molecule spec into dictionary from other schemas. |
|
Format a string representation of QM molecule. |
|
Forms molecular and fragment charge and multiplicity specification by completing and reconciling information from argument, supplemented by physical constraints and sensible defaults. |
qcelemental.molutil Package
Functions
|
Use Kabsch algorithm to find best alignment of geometry cgeom onto rgeom while sampling atom mappings restricted by runiq and cuniq. |
|
Generate a random or directed translation, rotation, and atom shuffling. |
|
Finds connected atoms based off of a covalent radii metric. |
|
Finds optimal translation and rotation to align cgeom onto rgeom via Kabsch algorithm by minimizing the norm of the residual, \(|| R - U * C ||\). |
|
Returns the molecular formula for a list of symbols. |
|
Reorders a molecular formula. |
qcelemental.testing Module
Functions
|
Returns True if two integers, strings, booleans, or integer arrays are element-wise equal. |
|
Function to compare Molecule dictionaries. |
|
Recursively compares nested structures such as dictionaries and lists. |
|
Returns True if two floats or float arrays are element-wise equal within a tolerance. |
qcelemental.models Package
Classes
Facilitates the application of the simple transformation operations defined by |
|
The MolSSI Quantum Chemistry Schema |
|
Results from a CMS program execution. |
|
Named properties of quantum chemistry computations following the MolSSI QCSchema. |
|
Old class for pydantic docstring before autodoc-pydantic came about. |
|
A quantum chemistry basis description. |
|
Complete description of the error from an unsuccessful program execution. |
|
|
Allowed computation driver values. |
Record indicating that a given operation (program, procedure, etc.) has failed and containing the reason and input data which generated the failure. |
|
The physical Cartesian representation of the molecular system. |
|
QCSchema input directive for geometry optimization. |
|
QCSchema results model for geometry optimization. |
|
QCSchema extension of pydantic.BaseModel. |
|
Provenance information. |
|
QC Result Schema. |
|
QC Input Schema. |
|
QC Result Properties Schema. |
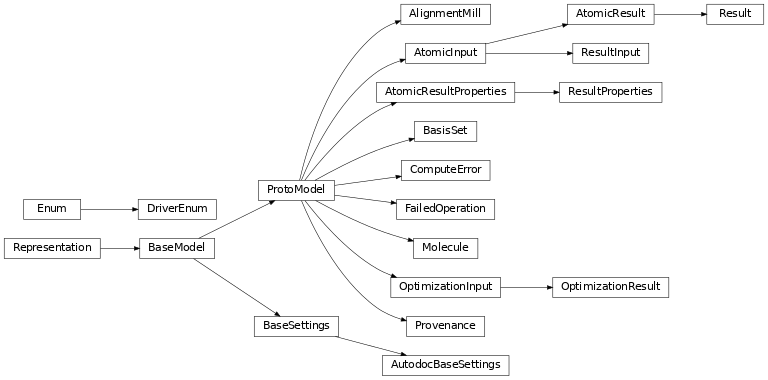
Class Inheritance Diagram